BaseVariableImportance#
- class hidimstat.base_variable_importance.BaseVariableImportance[source]#
Bases:
BaseEstimatorBase class for variable importance methods.
This class provides a foundation for implementing variable importance methods, including feature selection based on importance scores and p-values.
- Attributes:
- importances_array-like of shape (n_features,), default=None
The computed importance scores for each feature.
- pvalues_array-like of shape (n_features,), default=None
The computed p-values for each feature.
Methods
selection(k_best=None, percentile=None, threshold=None, threshold_pvalue=None)
Selects features based on importance scores and/or p-values using various criteria.
_check_importance()
Checks if importance scores and p-values have been computed.
- importance_selection(k_best=None, percentile=None, threshold_max=None, threshold_min=None)[source]#
Selects features based on variable importance.
- Parameters:
- k_bestint, default=None
Selects the top k features based on importance scores.
- percentilefloat, default=None
Selects features based on a specified percentile of importance scores.
- threshold_maxfloat, default=None
Selects features with importance scores below the specified maximum threshold.
- threshold_minfloat, default=None
Selects features with importance scores above the specified minimum threshold.
- Returns:
- selectionarray-like of shape (n_features,)
Binary array indicating the selected features.
- pvalue_selection(k_lowest=None, percentile=None, threshold_max=0.05, threshold_min=None, alternative_hypothesis=False)[source]#
Selects features based on p-values.
- Parameters:
- k_lowestint, default=None
Selects the k features with lowest p-values.
- percentilefloat, default=None
Selects features based on a specified percentile of p-values.
- threshold_maxfloat, default=0.05
Selects features with p-values below the specified maximum threshold (0 to 1).
- threshold_minfloat, default=None
Selects features with p-values above the specified minimum threshold (0 to 1).
- alternative_hypothesisbool, default=False
If True, selects based on 1-pvalues instead of p-values.
- Returns:
- selectionarray-like of shape (n_features,)
Binary array indicating the selected features (True for selected).
- fdr_selection(fdr, fdr_control='bhq', reshaping_function=None, two_tailed_test=False)[source]#
Performs feature selection based on False Discovery Rate (FDR) control.
- Parameters:
- fdrfloat
The target false discovery rate level (between 0 and 1)
- fdr_control: {‘bhq’, ‘bhy’}, default=’bhq’
The FDR control method to use: - ‘bhq’: Benjamini-Hochberg procedure - ‘bhy’: Benjamini-Hochberg-Yekutieli procedure
- reshaping_function: callable or None, default=None
Optional reshaping function for FDR control methods. If None, defaults to sum of reciprocals for ‘bhy’.
- two_tailed_test: bool, default=False
If True, performs two-tailed test selection using both p-values for positive effects and one-minus p-values for negative effects. The sign of the effect is determined from the sign of the importance scores.
- Returns:
- selectedndarray of int
Integer array indicating the selected features. 1 indicates selected features with positive effects, -1 indicates selected features with negative effects, 0 indicates non-selected features.
- Raises:
- ValueError
If importances_ haven’t been computed yet
- AssertionError
If pvalues_ are missing or fdr_control is invalid
- fwer_selection(fwer, procedure='bonferroni', n_tests=None, two_tailed_test=False)[source]#
Performs feature selection based on Family-Wise Error Rate (FWER) control.
- Parameters:
- fwerfloat
The target family-wise error rate level (between 0 and 1)
- procedure{‘bonferroni’}, default=’bonferroni’
The FWER control method to use: - ‘bonferroni’: Bonferroni correction
- n_testsint or None, default=None
Factor for multiple testing correction. If None, uses the number of clusters or the number of features in this order.
- two_tailed_testbool, default=False
If True, uses the sign of the importance scores to indicate whether the selected features have positive or negative effects.
- Returns:
- selectedndarray of int
Integer array indicating the selected features. 1 indicates selected features with positive effects, -1 indicates selected features with negative effects, 0 indicates non-selected features.
- plot_importance(ax=None, ascending=False, feature_names=None, **seaborn_barplot_kwargs)[source]#
Plot feature importances as a horizontal bar plot.
- Parameters:
- axmatplotlib.axes.Axes or None, (default=None)
Axes object to draw the plot onto, otherwise uses the current Axes.
- ascending: bool, default=False
Whether to sort features by ascending importance.
- **seaborn_barplot_kwargsadditional keyword arguments
Additional arguments passed to seaborn.barplot. https://seaborn.pydata.org/generated/seaborn.barplot.html
- Returns:
- axmatplotlib.axes.Axes
The Axes object with the plot.
- get_metadata_routing()[source]#
Get metadata routing of this object.
Please check User Guide on how the routing mechanism works.
- Returns:
- routingMetadataRequest
A
MetadataRequestencapsulating routing information.
- get_params(deep=True)[source]#
Get parameters for this estimator.
- Parameters:
- deepbool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
- Returns:
- paramsdict
Parameter names mapped to their values.
- set_params(**params)[source]#
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as
Pipeline). The latter have parameters of the form<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters:
- **paramsdict
Estimator parameters.
- Returns:
- selfestimator instance
Estimator instance.
Examples using hidimstat.base_variable_importance.BaseVariableImportance#
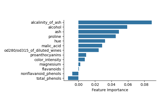
Conditional Feature Importance (CFI) on the wine dataset

Leave-One-Covariate-Out (LOCO) feature importance with different regression models
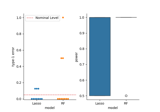

Distilled Conditional Randomization Test (dCRT) using Lasso vs Random Forest learners

Feature Importance on diabetes dataset using cross-validation

Conditional vs Marginal Importance on the XOR dataset
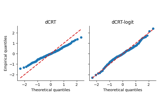

Conditional Randomization Test for Sparse Logistic Regression
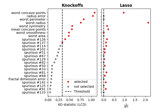
Controlled multiple variable selection on the Wisconsin breast cancer dataset
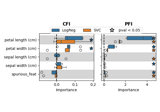

Measuring Individual and Group Variable Importance for Classification

Pitfalls of Permutation Feature Importance (PFI) on the California Housing Dataset